Maternal postpartum sedentary life exacerbates the metabolic impact of moderately fat-rich diet in male mice
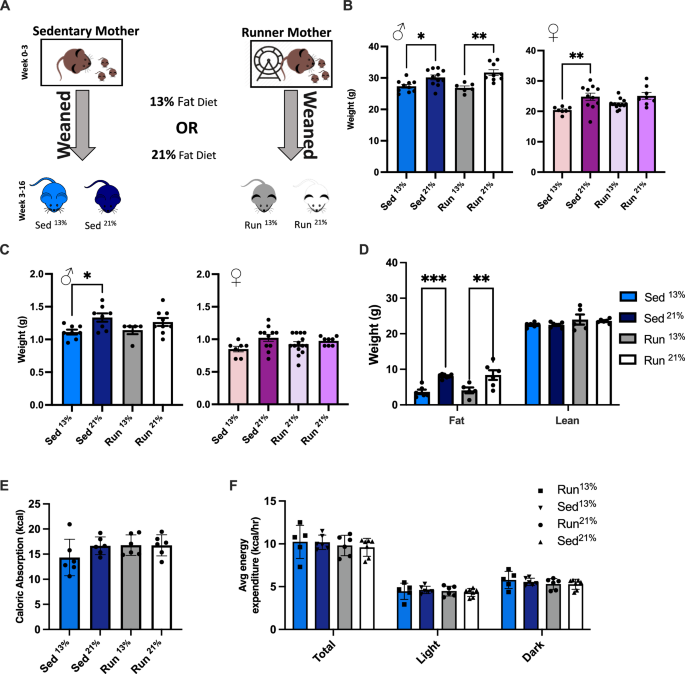
Experimental design: metabolic consequences of maternal postpartum physical activity and postweaning offspring diet
We tested the effect of maternal postpartum physical activity or the lack thereof between P2 and P21 in combination with the offspring’s diet from weaning at P21 to adulthood on the metabolic health of the adult offspring (Fig. 1A). The offspring diet was either standard, containing 13% fat calories (Picolab 5053) or 21% fat calories (Picolab 5058).
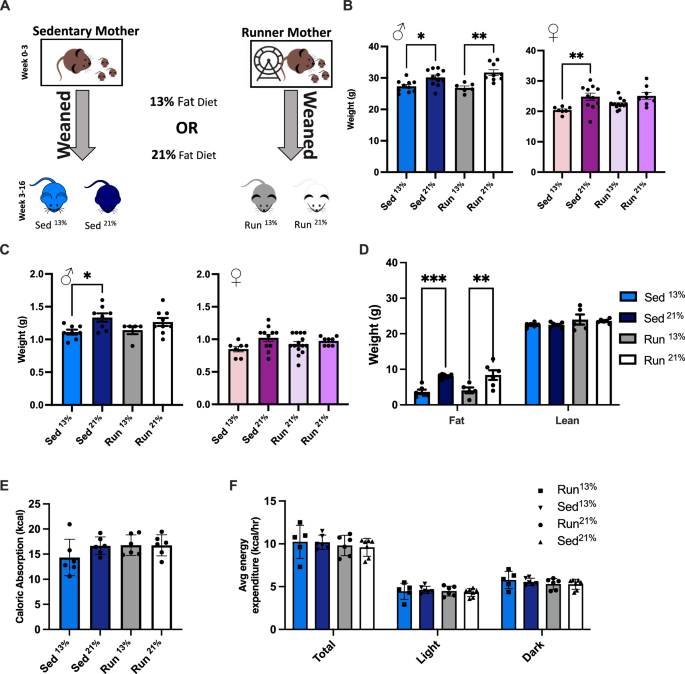
A Experimental design of generating four groups of offspring. Sed offspring were raised by mothers housed in standard cages while Run offspring were raised by mothers from P2 in cages equipped with running wheels. At P21, male and female offspring were transferred to standard cages on either 13% or 21% fat calorie diet. B Sed and Run male offspring on 13% fat calorie diet had comparable weights. The 21% diet, relative to the 13% diet, resulted in a modest weight gain in both Sed and Run males. One-way ANOVA, F(3,32) = 8.354, P = 0.0003. Sed13% vs. Sed21%, Tukey’s test for multiple comparisons, *p = 0.0410; Run13% vs. Run21%, **p = 0.0021. N = 10 (Sed13%), 11 (Sed21%), 6 (Run13%), 9 (Run21%). Female offspring showed a similar trend, but weight gain was significant only in the Sed group. One-way ANOVA, F(3,36) = 5.540, P = 0.00031. Sed13% vs. Sed21%, Tukey’s test for multiple comparisons, p = 0.0088. N = 7 (Sed13%), 11 (Sed21%), 14 (Run13%), 8 (Run21%). C Increased liver weight in Sed21% males (left) but not females (right). Males, one-way ANOVA, F(3,26) = 3.422, P = 0.0319. Sed13% vs. Sed21%, Tukey’s test for multiple comparisons, *p = 0.0394. N = 8 (Sed13%), 8 (Sed21%), 5 (Run13%), 9 (Run21%). Females, one-way ANOVA, F(3,36) = 2.396, P = 0.0842. N = 7 (Sed13%), 11 (Sed21%), 14 (Run13%), 8 (Run21%). D Weight gain in Sed21% and Run21% males was due to increased body fat accumulation as measured by MRI. Two-way ANOVA, interaction between body composition and diet F(3,40) = 6.688, P = 0.0009; body composition F(1,40) = 1068, P < 0.0001; diet F(3,40) = 6.204, P = 0.0015. Sed13% vs. Sed21%, Tukey’s correction for multiple comparison, ****p = 0.0004; Run13% vs. Run21%, ***p = 0.0017. N = 6 (Sed13%), 7 (Sed21%), 5 (Run13%), 6 (Run21%). E, F Calorie absorption and expenditure were not influenced by postpartum maternal physical activity or postweaning offspring diet. Calories absorbed, One-way ANOVA, F(3,20) = 1.380, P = 0.2777. Expenditure, Two-way ANOVA, interaction between time and group, F(6,40) = 0.7514, P = 0.6120. N = 6 (Sed13%), 7 (Sed21%), 5 (Run13%), 6 (Run21%).
To this end, mothers with their newborn pups were randomly divided at P2 into 2 groups, both on the 13% diet. One group was housed in standard “sedentary” cages while the other was housed in cages equipped with running wheels (Sed and Run offspring, respectively). Litter size was limited to no more than six. As we reported earlier, sedentary and running mothers spent a similar amount of time caring for their pups during the dark and light periods19. Further, weight gain and survival of pups at weaning were indistinguishable between the two groups19.
At P21, Sed and Run offspring were weaned to standard cages (without running wheels) in groups of 4–5 and were randomly assigned to the 13% or 21% fat calorie diets until adulthood (12–16 weeks of age), after which they were subjected to behavioral and metabolic testing or used for tissue collection (Fig. 1A). Mothers were used only once in running experiments and were sacked afterwards to avoid the effect of repeated running on offspring behavior. The 13% diet is considered a normal fat diet22, while the 21% diet is a medium fat diet because its fat calorie content is between a normal and high fat diet. Although the medium-fat diet is similar to a Western diet, it has no detrimental effect if given to adult mice. In fact, it was designed to meet the energy needs of breeding mouse colonies, mice exposed to higher stress levels, and transgenic strains ( We opted not to use a fat (i.e., diet-induced obesity or DIO) diet because its 45–60% fat calorie content, although consistently produces metabolic disease characterized by obesity, dyslipidemia, insulin resistance, hypertension, and non-alcoholic fatty liver23,24, represents a dietary extreme that does not reflect common human dietary habits. The 21% fat calorie diet has a similar amount of protein as the 13% diet but contains proportionally less carbohydrates. We started the 21% fat calorie diet at weaning because it is now recognized that diet-induced obesity often begins in childhood25. Since mothers were always on the 13% diet, this design allowed us to study the interaction between maternal postpartum physical activity and offspring diet without the possible confounding effect of the 21% diet on maternal physical activity. Overall, this design generated four groups of offspring: Sed13%, Sed21%, Run13%, and Run21%.
Medium fat calorie diet increases both the body and liver weight in Sed males
We previously reported that the Run and Sed offspring had comparable weight at weaning (at 3 weeks of age) and that they were indistinguishable in weight gain on a normal 13% fat calorie diet during development, adult weight, adult total body composition, and adult metabolic parameters that included heat generation, energy expenditure, O2 consumption, and CO2 production19. Here, we asked if Sed and Run offspring respond differently to increased dietary fat. A moderate increase in fat calories from 13% to 21% in the diet from weaning resulted in increased adult weight in males (Fig. 1B). However, adult weight (on the 21% diet) was comparable in the Run and Sed males, indicating no apparent effect of maternal postpartum physical activity. Females showed a similar trend, but weight gain reached significance only in the Sed group (Fig. 1B).
Although body weight increased similarly in Sed and Run males on increased fat diet (Fig. 1B), Sed males on 21% medium fat diet, as compared Sed males on 13% normal fat diet, had increased liver weight (Fig. 1C). These data indicate that even a moderate increase in dietary fat from weaning at P21 results in liver abnormality and that this effect is neutralized by maternal postpartum physical exercise. In contrast to males, Sed females on 21% diet showed no significant increase in liver weight (Fig. 1C), indicating that females are not sensitive to a moderate increase in dietary fat calories. Liver represented only ~4% of the total body weight, explaining why the ~15% increase in liver weight in Sed21% males did not add significantly to their total weight when compared with the total weight of Run21% males (in Fig. 1B).
Since Sed males but not Sed females exhibited a gain in liver weight on a 21% fat calorie diet, only males were studied subsequently. The weight gain on the 21% fat calorie diet from weaning was due to increased fat accumulation (as measured by MRI) in both Sed and Run males (Fig. 1D). However, total caloric absorption from the GI track and energy expenditure of males were not altered by postpartum maternal activity and postweaning diet (Fig. 1E, F). Further, food consumption, respiratory exchange rate (RER), and locomotor activity through the day were unchanged across the 4 groups of males (Supplementary Fig. 1A–C). Therefore, the difference in liver weight between Sed21% and Run21% males was not due to differences in the amount of food consumed, calories absorbed or expended, or physical activity.
Finally, fasting (6 h) plasma triglyceride (TG) and free FA levels were not significantly changed by maternal running, 21% diet, and their combination, relative to levels in Sed mice on the 13% diet, although there was a trend toward reduced TG levels in Run as compared to Sed males on the 21% calorie diet (Supplementary Fig. 2A, B). The 21% fat calorie diet was not associated with increased fasting glucose level, and the glucose tolerance test showed no group difference at any time point, although there was an interaction between time and group (Supplementary Fig. 2C, ANOVA with Tukey post hoc.). Overall, both Sed and Run males on the 21% fat calorie diet had a largely normal glucose homeostasis, unlike the higher fat DIO diet that produces hyperglycemia and glucose intolerance26.
The 21% fat calorie diet induces lipid accumulation in the liver of Sed but not Run males
Consistent with their increased liver weight, Sed21% males exhibited lipid accumulation in the liver27, while the liver of Run21% males had normal weight and very few lipid droplets, like the liver of Sed and Run males on 13% fat calorie diet (Fig. 2A, B). Quantification revealed an ~ 3-fold increase in lipid droplets in Sed21% livers, compared to all other groups (Fig. 2C). Overall, these data indicate that even a moderate increase in fat calorie intake from weaning age can cause lipid accumulation in the liver of male C57BL/6 mice maintained under standard laboratory conditions (i.e., Sed21%), and that maternal exercise during the postnatal period can completely prevent the medium fat diet-induced liver pathology in the otherwise sedentary male offspring (i.e., Run21%).
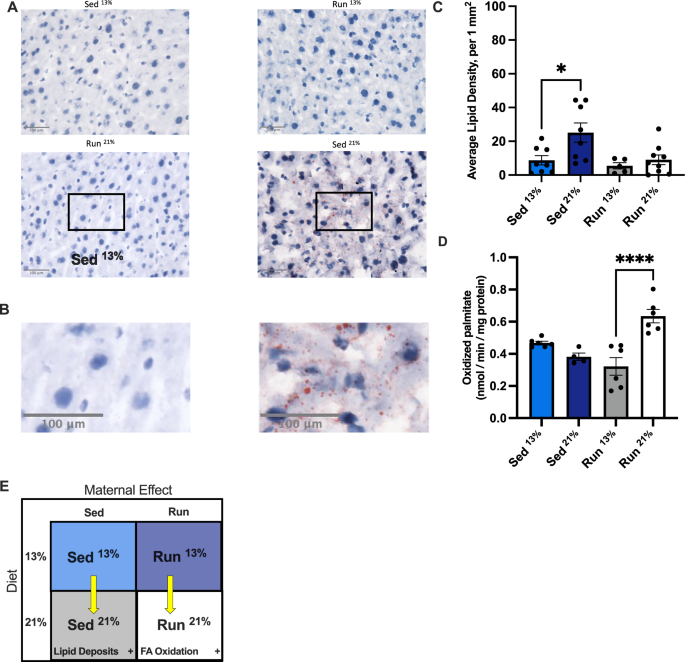
A Representative ×40 images of oil red O-stained liver sections show lipid deposits in Sed21% males but not in Sed13%, Run21%, and Run13% males. B Higher magnification images of Sed21% and Run21% livers highlighting the difference in lipid deposits. C Increased lipid droplet density in the liver of Sed21% males. One-way ANOVA, F(3,26) = 5.198, P = 0.0060. Sed13% vs. Sed21%, Tukey’s test for multiple comparisons, *p = 0.0224. N = 8 (Sed13%), 8 (Sed21%), 5 (Run13%), 9 (Run21%). D Increased beta oxidation in Run22% male livers. One-way ANOVA, F(3,18) = 12.87, P < 0.0001. Run13% vs. Run21%, Tukey’s test for multiple comparisons, *p < 0.0001. N = 6 (Sed13%), 4 (Sed21%), 6 (Run13%), 6 (Run21%). E Summary of liver and brown adipose tissue changes by postweaning 21% fat diet and the interaction between diet and preweaning maternal environment.
Increased β-oxidation in the liver of Run offspring on a medium-fat diet
We reasoned that the lack of lipid accumulation in the Run offspring on the 21% fat diet was due to metabolic adaptive plasticity, induced by the preweaning Run maternal environment, that allowed coping with a moderately high-fat diet after weaning. To test this notion, we measured FA oxidation from liver extracts from all four groups of males (Fig. 2D). FA oxidation in Sed mice on the 21% and 13% diets was comparable, in agreement with reports showing no or variable effects of even higher (40–60%) fat calorie diet on FA oxidation and respiration in C56BL sedentary mice28,29,30,31. However, Run mice on the 21%, relative to the 13% fat calorie diet, had increased FA oxidation, indicating that the combined effects of the preweaning Run maternal environment and postweaning 21% diet increased the liver’s capacity for FA oxidation. Overall, the liver showed opposite changes on 21% medium relative to 13% normal fat diet in Sed and Run mice: increased lipid deposits with no change in FA oxidation in Sed and increased FA oxidation with no lipid accumulation in Run mice (Fig. 2E).
Lack of liver lipid accumulation in Run males on a medium-fat diet is associated with metabolic adaptation in the liver
Next, we sought to determine factors that either contribute to increased lipid accumulation in the liver of Sed21% mice or prevent the development of this phenotype in Run21% mice. LC/MS-based metabolite profiling identified 30 FA species, including even and odd chain and hydroxyl and keto forms. The sum of the z-scored ion abundances of these species indicated a significant diet-induced increase but no running maternal effect (Supplementary Fig. 3, Supplementary Table 1). However, analysis of individual FA species (Fig. 3A) revealed not only a diet-induced effect (Sed21% vs. Sed13% and Run21% vs. Run13%) but also a Run maternal effect (Run13% vs. Sed13%), and interaction between the two (Run21% vs. Sed13%) (Fig. 3B, left panel). Run21% livers contained less medium-chain FA (MCFA) species (C6 and C8 carbons in aliphatic tail) than Sed21% and even Sed13% livers, indicating a main effect of the Run maternal environment (Fig. 3B, right panel, caproic acid and alpha-lipoic acid). Reduced levels of these MCFAs in Run21% mice may suggest that less is available for TG synthesis and lipid accumulation.
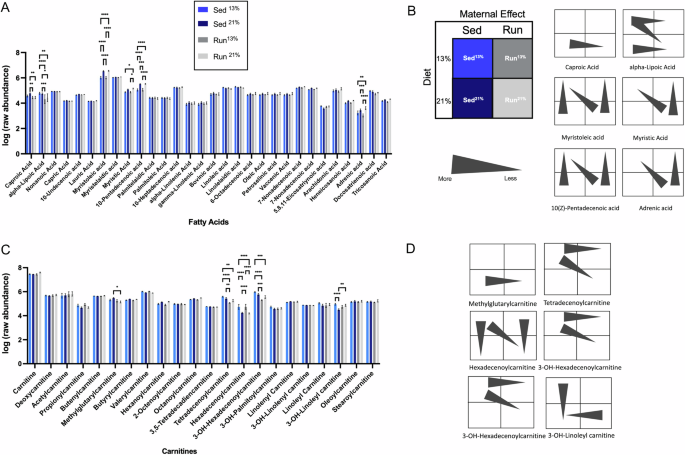
A FA levels in Sed13%, Sed21%, Run13%, and Run21% livers. Two-way ANOVA; interaction between groups and metabolites, F(87,629) = 2.064, *p < 0.0001; group effect, F(3,629) = 20.58, *p < 0.0001. Tukey’s test for multiple comparisons. C6, caproic acid, Run21% vs. Sed21%, **p = 0.0070; Run13% vs. Sed21%, **p = 0.0078. C8, alpha-lipoic acid, Run13% vs. Sed13%, ****p < 0.0001; Run13% vs. Sed21%, ****p = 0.0002; Run21% vs. Sed13%, ****p = 0.0030; Run21% vs. Sed21%, *p = 0.0160. C14, myristoleic acid, Run21% vs. Run13%, ****p < 0.0001; Run13% vs. Sed21%, ****p < 0.0001; Run21% vs. Sed13%, ****p < 0.0001; Sed21% vs Sed13%, ****p < 0.0001. C14, myristic acid, Run21% vs. Run13%, *p = 0.0285; Run21% vs. Sed13%, *p = 0.0189. C15:1, 10-pentadecenoic acid, Run21% vs. Run13%, ****p < 0.0001; Run13% vs Sed21%, ***p = 0.0001; Run21% vs. Sed13%, ****p < 0.0001; Sed21% vs. Sed13%, ****p < 0.0001. C22, adrenic acid, Run21% vs. Run13%, ****p < 0.0001; Run13% vs. Sed21%, ***p = 0.0004; Run21% vs. Sed13%, **p = 0.0015. N = 7, Sed13%; N = 6, Sed21%; N = 6, Run13%; N = 6, Run21%. B Schematic representation of main trends and directions of significant changes in (A) in pairwise comparisons between groups, indicating the effect of diet (vertical arrowheads), effect of the maternal environment (horizontal arrowheads), and the combined effect of diet and maternal environment (diagonal arrowheads). While medium-chain FAs (C6 and C8) are reduced, long-chain FAs are increased in Run21% relative to Sed13% and Run13% mice. C Carnitine (C0) and acylcarnitine (C2-C18) levels in Sed13%, Sed21%, Run13%, and Run21% livers. Two-way ANOVA; interaction between groups and metabolite, F(63,461) = 2.756, p < 0.0001, group effect, F(3,461) = 5.170, p = 0016. Tukey’s test for multiple comparisons. C5, methylglutarylcarnitine, Run21% vs. Sed21%, *p = 0.0148. C14, tetradecenoylcarnitine, Run13% vs. Sed13%, ****p < 0.0001; Run21% vs. Sed13%, **p = 0.0072; Run13% vs Sed21%, **p = 0.0061. C16, hexadecenoylcarnitine, Sed21% vs. Sed13%, ****p < 0.0001; Run21% vs. Sed13%, ****p < 0.0001; Run13% vs. Sed21%, ****p < 0.0001; Run21% vs. Run13%, ****p < 0.0001. C16, 3-OH-Hexadecenoylcarnitine, Run13% vs. Sed13%, ****p < 0.0001; Run21% vs. Sed13%, ***p = 0.0006; Run13% vs Sed21%, ***p = 0.0001. C18, 3-OH-linoleyl carnitine, Sed21% vs. Sed13%, ****p < 0.0001; Run21% vs. Sed21%, **p = 0.0044. N = 7, Sed13%; N = 6, Sed21%; N = 6, Run13%; N = 6, Run21%. D Schematic representation of main trends and directions of significant changes in long-chain acylcarnitine species in (C) showing an overall reduction of acylcarnitines in Run21% liver.
In contrast, four long-chain FAs (LCFAs) had significantly higher levels in Run21% than in Run13% and Sed13% livers (myristoleic acid, C14:1; myristic acid, C14:0; pentadecenoic acid, 15:1; and adrenic acid, C22:4), two of which (myristic acid and adrenic acid) were not increased in Sed21% mice (relative to Sed13% mice) indicating more diet-induced FAs in Run than in Sed mice. In contrast to MCFAs, transport of LCFAs into the mitochondria for β-oxidation requires carnitines32; thus, the higher LCFA levels in Run21% livers, given increased FA β-oxidation (Fig. 2D), may indicate alterations in the availability of carnitine and/or saturation of the activity of the carnitine cycle.
Carnitine forms acylcarnitines with LCFAs through conjugation for transport across the mitochondrial membrane33. Carnitine level was not changed by either the maternal effect or diet (Fig. 3C), but a number of acylcarnitine species were reduced by the Run maternal effect or by the combined effect of maternal environment and diet (C5 methylglutarylcarnitine, C14:1 tetradecenoylcarnitine, C16:1 hexadecenoylcarnitine, and C16:1-OH 3-OH-hexadecenoylcarnitine), resulting in an overall reduction of these acylcarnitine species in Run21% livers relative to Sed21%, Sed13%, or Run13% livers (Fig. 3C, D). Reduced acylcarnitines, given the unchanged carnitine level, implies increased flux of FAs to β-oxidation in Run21% mice, which is consistent with increased liver FA oxidation rates in these mice (Fig. 2D), and may explain, at least partly, their resistance to liver lipid accumulation on the 21% fat calorie diet (Fig. 2E). The only exception was a long-chain C18 acylcarnitine (3-OH linoleyl carnitine) whose level was increased in Run21% relative to Sed21% mice.
Single-nucleus transcriptomic analysis of liver cells
We reasoned that differences in liver lipid accumulation, FA oxidation, and FA metabolite levels between the Sed and Run offspring on the 21% fat calorie diet should be reflected in differences in lipid catabolism and/or biosynthesis-related gene expression in the liver. Further, hepatic immune cells may also exhibit changes in gene expression because activation of local Kupffer cells and stellate cells may accompany lipid accumulation in Sed21% liver34,35. To simultaneously profile both hepatocytes and various types of non-parenchymal cells from liver tissue, including Kupffer cells and stellate cells, we performed single nucleus (sn)RNA-Seq using the 10x sequencing platform36,37,38,39. Liver samples were pooled from three individual male offspring per group, each from a separate mother.
A total of 14,136, 16,891, 11,594, and 14,570 cells from Sed13%, Sed21%, Run13%, and Run21% livers, respectively, passed quality control and were visualized as clusters by Uniform Manifold Approximation and Projection (UMAP). The overall distribution of cell clusters showed only minor differences, mainly in the hepatocyte clusters, across the four groups of mice (Supplementary Fig. 4). Therefore, we produced a combined UMAP profile to assign clusters to cell types, using cell type specific markers reported earlier in the mouse liver38 (Fig. 4, and Supplementary Fig. 5 and Supplementary Table 2 for cluster markers). Based on cell type-specific markers, we identified seven hepatocyte, two Kupffer cell, and four stellate cell clusters (Fig. 4A–D). Seven clusters formed a mixed population of various endothelial cells (Fig. 4E), and additional clusters were represented by B cells and NK cells (Fig. 4F, G).
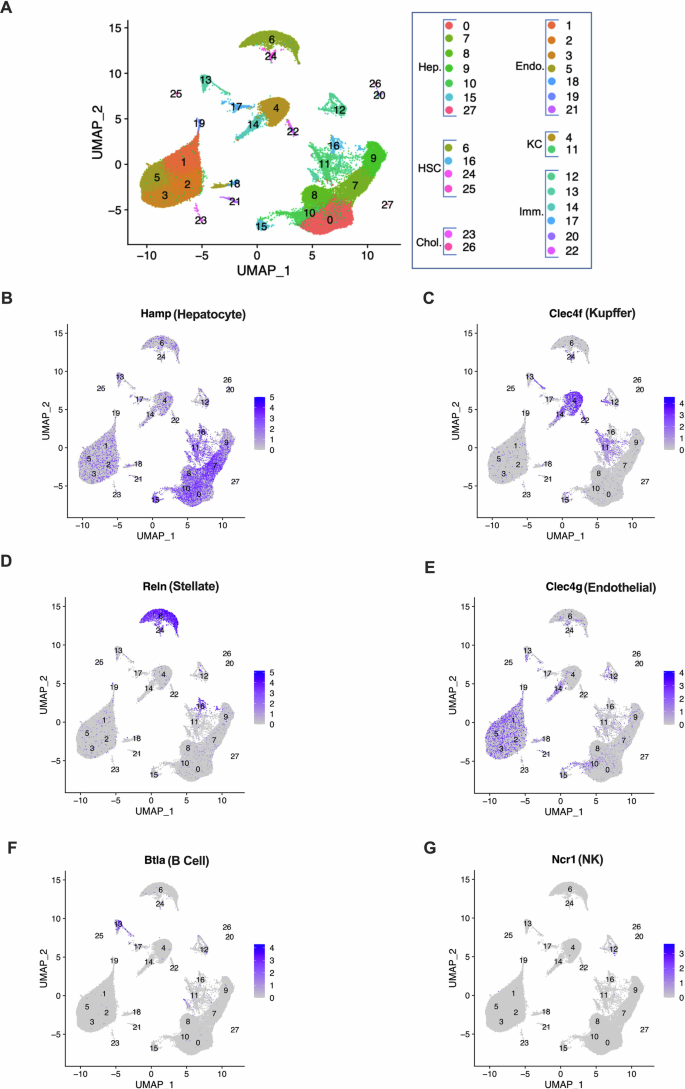
A Combined UMAP plot of transcriptional profiles of single nuclei from Run21%, Run13%, Sed21%, and Sed13% livers. All identified hepatocytes (Hep) and non-parenchymal cells, including Kupfer cells (KC), hepatic stellate cells (HSC), endothelial cells (Endo), cholangiocytes (Chol), and immune cells (Imm) are shown. B–G Expression of cluster marker genes specific for the major cell types in the liver. Hamp, hepatocytes; Clec4, Kupffer cells; Reln, Stellate cells; Clec4g, Endothelial cells; Btla, B cells; Nrc1, NK cells.
Medium-fat diet is associated with more transcriptional changes in the liver of Sed than Run mice
Next, we sought to identify the effects of diet, maternal environment, and the combination of the two on the transcriptomic profile of hepatocytes and other liver cells using snRNA-Seq data from the four groups of mice (Fig. 5A, E, I, red vertical, diagonal, and horizontal arrows, respectively).
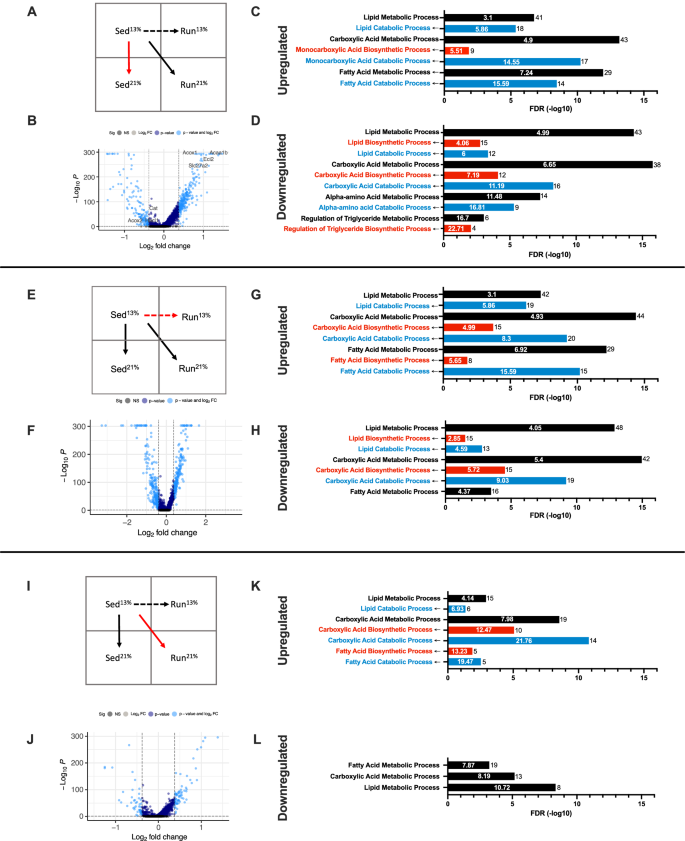
A Diagram of transcriptomic comparisons, red arrow indicating the Sed21% vs. Sed13% comparison. B 21% fat calorie diet-induced up- (+) and downregulated (−) genes in Sed mice indicated by light blue dots (FC > 1.3-fold, log10P < 0.05). Purple dots, log10P < 0.05; light gray, FC > 1.3; dark gray, not significant. C, D Top gene ontology (GO) biological processes (nonredundant categories with <1000 genes), overrepresented in Sed21% vs. Sed13% DEGs. Both upregulated (C) and downregulated (D) DEGs are enriched in lipid, carboxylic acid, and amino acid metabolism-related GO biological processes (black bars). Splitting DEGs to catabolic (blue) and biosynthetic (red) genes shows that both up- and downregulated genes are more enriched in catabolic processes. E Transcriptomic comparison of Run13% and Sed13% hepatocytes (red arrow). F Maternal environment-induced DEGs. G, H Up- and downregulated DEGs are also enriched in lipid/carboxylic acid catabolic (blue) and biosynthetic processes (red). I Comparison of Run21% vs. Sed13% transcriptomes (red arrow). J Fewer DEGs are identified in Run than in Sed (see B) mice on the 21% diet. K, L Significance of enrichment of biological processes is also mitigated in the Run offspring, but up- and especially downregulated genes are still mostly enriched in catabolic (blue) processes as compared to biosynthetic (red) processes.
By using the combined set of hepatocyte clusters o, 7, 8, 9, 10, 15, and 27 (Fig. 4A, B), we identified 424 diet-related (red arrow in Fig. 5A) differentially expressed genes (DEGs, >1.3-fold, adj. p < 0.05) between Sed21% and Sed13% mice (Fig. 5B). The 265 upregulated genes (Supplementary Table 3) were highly enriched in the overlapping Gene Ontology biological functions (GO40, nonredundant, <1000 genes) of “lipid and carboxylic acid metabolic processes” (Fig. 5C, black columns) that, on further dissection, were found to be dominated, based on lower FDRs, by catabolic processes (lipid, carboxylic acid, and FA), with much lower representation by biosynthetic processes (Fig. 5C, blue and red columns, respectively, see also corresponding genes in Fig. 6A). Upregulated genes in these GO categories included Acox1 encoding an acyl-coenzyme A oxidase involved in the first step of peroxisomal FA oxidation (i.e., desaturation of acyl-CoAs to 2-trans-enoyl-CoAs), Slc27a2 whose protein product transports exogenous long-chain fatty acids (FA) and activates very-long-chain FAs for degradation, Eci2 (enoyl-CoA delta isomerase 2) involved in the oxidation of unsaturated FAs in mitochondria, and Acaa1b (3- ketoacyl-CoA thiolase B) involved in peroxisomal FA oxidation. These changes suggest a compensatory transcriptomic response of Sed21% livers to increased dietary fat calories. However, the 159 downregulated genes in Sed21% vs. Sed13% livers were also enriched in the same biological processes (Figs. 5D and 6A). This group of downregulated genes included Acox2, Cat (carnitine O-acetyltransferase) that catalyzes the production of acylcarnitines, and Hacl1 (2-hydroxyacyl-CoA lyase 1) involved in the degradation of methyl-branched FAs and 2-hydroxy long-chain FAs (Fig. 6A, Supplementary Table 3). Downregulated Sed21% vs. Sed13% genes were also enriched in amino acid metabolic processes as part of the broader carboxylic acid catabolic pathway and were again biased toward catabolic over biosynthetic processes (Fig. 5D). However, we detected no changes in liver amino acids by LC-MS (not shown). Overall, the transcriptional response of hepatocytes to increased fat calories in Sed mice was preferentially associated with lipid/FA catabolic processes, involving both upregulated and less prominently downregulated genes. The opposite regulation of some genes within the same biological processes (even in the same gene family; i.e., Acox1 and Acox2), specifically in FA oxidation, and the unaltered activity of liver β oxidation in Sed21% mice, suggest that the lipid/FA-related transcriptome is reorganized but does not effectively compensate for the increased fat calorie intake in SED mice. Interestingly, two of the downregulated genes, Acox2 and Hacl1, were upregulated on the 21% fat calorie diet in Run mice that exhibited increased β oxidation and no lipid accumulation in the liver (Fig. 6A; Run21% vs. Run13% DEGs in parentheses).
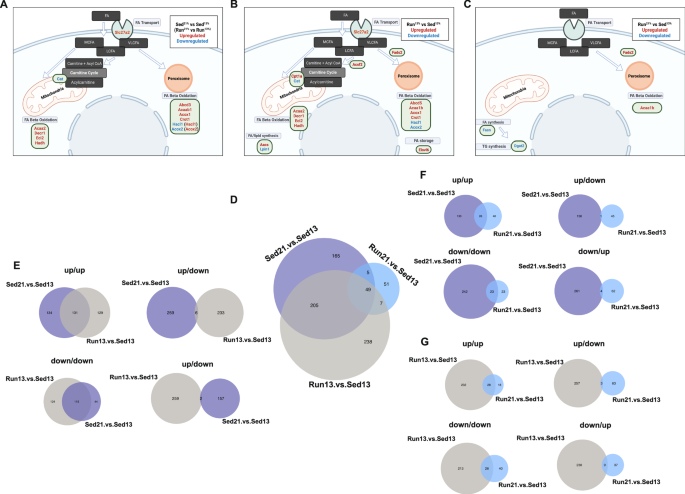
A Sed21% vs. Sed13% DEGs enriched in lipid metabolic processes mapped to FA transport and beta oxidation metabolic pathways. DEGs in Run21% vs. Run13% are in parentheses. Abcd3 (ATP binding cassette subfamily D member 3), Acaa1b (3- ketoacyl-CoA thiolase B), Acox1 and Acox2 (acyl-coenzyme A oxidase), Cat (carnitine O-acetyltransferase), Crot (carnitine O-octanoyltransferase), Eci2 (enoyl-CoA delta isomerase 2), Decr1 (2,4-dienoyl-CoA reductase 1), Hacl1 (2-hydroxyacyl-CoA lyase 1), Hadh (3-hydroxyacyl-CoA dehydrogenase), Slc27a2 (solute carrier for long-chain FAs. FA: fatty acid; MFA, medium-chain FA; LCFA, long-chain FA; VLCA, very-long-chain FA. B Run13% vs. Sed13% DEGs mapped to lipid/FA metabolic processes. Hacd3 (3-Hydroxyacyl-CoA dehydratase 3), Cpt1a (carnitine palmitoyltransferase 1A), Acsf2 (acyl-CoA synthetase family member 2). C Run21% vs. Sed13% DEGs mapped to lipid/FA metabolic processes. Fasn (FA synthase), Dgat2 (diacylglycerol acyltransferase). D Overlap between diet, maternal environment, and diet + maternal environment-induced DEGs. E–G Overlap between DEGs related to pairs of conditions in (D) is split into up- and downregulated genes.
The Run maternal environment alone (i.e., Run13% vs Sed13%, Fig. 5E, horizontal arrow) produced 266 up- and 233 downregulated DEGs (Fig. 5F, Supplementary Table 3) that were also enriched in “carboxylic acid and lipid/FA metabolic processes” with a bias toward catabolic processes (Fig. 5G, H, Supplementary Table 3). Remarkably, many of the genes upregulated by the 21% diet in Sed mice were also upregulated by the postpartum maternal environment in the offspring on the 13% diet including Slc27a2, Acaa2, Decr1, Eci2, Hadh, Acaa1b, Acox1, and Crot (Fig. 6B). Additional upregulated genes included Cpt1a, encoding carnitine palmitoyltransferase 1 A that catalyzes the conjugation of FAs with carnitine to acylcarnitines. Other upregulated genes that may also promote FA catabolism and/or reduce lipid accumulation were Acss2 (encoding an Acyl-CoA Synthetase), Fads1 (fatty acid desaturase) involved in desaturation and elongation of polyunsaturated fatty acids41 (PUFAs), and Abca8a (an ABC transporter) promoting cholesterol efflux42. However, upregulated genes also included some with potential lipogenic effect, such as Aacs (acetoacetyl-CoA synthetase) that promotes the synthesis of cholesterol and FAs from ketone bodies and Elovl6 involved in FA elongation and lipid accumulation in the liver43,44. Some of the downregulated genes in Run13% mice were also downregulated in the liver of Sed21% mice, such as Acox2, Hacl1, and Cat, again suggesting a similar transcriptomic response to two different environmental effects. Finally, Lpin1, encoding phosphatidate phosphatase that converts phosphatidate to diacylglycerol to generate TAG for accumulation in the liver as lipid droplets45 was also downregulated. Overall, the Run maternal environment produced a liver transcriptional profile consistent with improved lipid/FA homeostasis in the otherwise sedentary offspring46.
Notably, transcriptomic response of Run mice to the 21% fat diet (representing the combined effect of fat diet and Run maternal environment), relative to that of Sed13% mice (Fig. 5I, diagonal red arrow) was modest with only 66 upregulated and 46 downregulated DEGs (Fig. 5J), although also involved “lipid catabolic and biosynthetic processes”, though with relatively low enrichment scores (Fig. 5K, L, Supplementary Table 3). Fads2 and Acaa1b, which were both upregulated in Sed21% and Run13% mice were also upregulated in Run21% mice (all relative to Sed13% mice, Fig. 6C). Specific for Run21% were the downregulation of Fasn, encoding FA synthase, and Dgat2, encoding diacylglycerol acyltransferase 2 that directs the de novo synthesized FA to TG synthesis and storage in lipid droplets. The changes in Fasn and Dgat2 could potentially reduce de novo FA and TG synthesis, respectively, and contribute to the resistance of the liver of Run mice to lipid accumulation. Finally, despite the increase in liver β oxidation in Run21% mice, corresponding genes that were DEGs in Sed21% vs Sed13% and Run13% vs Sed13% mice (Fig. 6A, B) were not upregulated in Run21% vs. Sed13% hepatocytes (Fig. 6C) indicating that increased β oxidation of the Run21% offspring may primarily be due to a metabolic rather than transcriptional adaptation. This notion was further supported by a minimal transcriptional change in nuclear encoded mitochondrial genes in Run21% (vs. Sed13%) hepatocytes (only one DEG), while genes encoding mitochondrial matrix-, transmembrane-, carrier-, and transporter-proteins, as well as mitochondrial enzymes, were differentially expressed by the diet and Run maternal environment (Sed21% vs Sed13% and Run13% vs Sed13%, Supplementary Fig. 6). Overall, DEGs associated with the combination of the Run maternal environment and 21% fat diet suggests a moderate transcriptional adaptation in FA and TG catabolism and synthesis, and in mitochondrial functions generally, presumably because of the relatively undisturbed lipid/FA homeostasis with increased activity in mitochondrial FA oxidation.
Finally, direct comparison of Run and Sed males on the 21% fat calorie diet again showed a relatively large number of Run21% vs. Sed21% DEGs in the liver (141 up- and 183 downregulated, respectively, Supplementary Fig. 7A), with a particularly high enrichment of upregulated genes in “carboxylic acid catabolic processes” (Supplementary Fig. 7B), consistent with the effect of the Run maternal environment on the liver transcriptome, whether combined with 13% (Fig. 5G, H) or 21% diet (Supplementary Fig. 7B). Given the lack of lipid accumulation and modest transcriptomic response of Run21% liver (relative to Sed13% controls), many of the Run21% vs. Sed21% DEGs were due to the significant changes in Sed21% livers and thus were in the opposite direction (Eic2, Slc27a2, Acox1, Acaa1b; Acox2, and Hacl1). It follows that some of the other Run21% vs. Sed21% DEGs that were also Run21% vs. Sed13% DEGs had the same directionality of change (Dgat2, Fasn, Lpin1, and Abhd2).
Medium-fat diet and maternal running produce partially overlapping transcriptional changes in the liver
DEGs associated with increased fat calorie diet (Sed21% vs. Sed13%) and maternal postpartum physical activity (Run13% vs. Sed13%) were not only enriched in the same GO functional annotations (Fig. 5C, D, G, H), but many DEGs within these functional categories were the same genes with the same directionality of change (Fig. 6A, B). This indicates that the two very different environmental influences (offspring postweaning exposure to medium fat diet and the preweaning effect of maternal exercise) alter the transcription many of the same genes (Fig. 6D). Indeed, approximately half of the Sed21% vs. Sed13% and Run13% vs. Sed13%DEGs were shared (Fig. 6D), 97.6% of which were up or downregulated in both conditions (131 up- and 115 downregulated in both conditions vs. 6 and 2 that changed in the opposite direction, Fig. 6E) suggesting that transcriptional changes in moderate fat may represent adaptive changes, similar to those induced by maternal exercise on the offspring liver. Similarly, approximately half of the Run21% vs. Sed13% DEGs (representing the combined effect of fat diet and maternal exercise), were also present in the larger sets of diet (Sed21% vs. Sed13%) and Run maternal effect (Run21% vs. Sed13%) DEGs, with 44% of them overlapping with both the diet and maternal effect DEGs (Fig. 6D), of which 90.7% and 78.5% of shared DEGs had the same directionality of transcriptional change, respectively (Fig. 6F, G). Taken together, approximately half of the DEGs induced by the 21% diet in Sed mice, the running maternal environment (on the 13% diet), and the combination of the two (21% diet and running maternal effect) were shared, an unexpected finding given the fundamental differences between these conditions. This suggests a transcriptional response to modify, presumably improve, lipid and FA homeostasis not only in the Run maternal group but also in the Sed group on the 21% fat calorie diet.
Transcriptional profile of Kupffer and stellate cells indicates a moderate liver injury in metabolically compromised Sed21% mice
Next, we analyzed the transcriptomic response of non-parenchymal cells in the liver to the 21% fat calorie diet in Sed and Run mice. Kupffer cells are resident liver macrophages47 and a critical component of the phagocytic system in the liver48. Kupffer cells can exhibit an activated/proinflammatory state characterized by IL-6 and TNF expression, and Kupffer cells seem to be directly involved in high-fat diet-induced hepatic steatosis49,50,51. We found 190 up- and 326 downregulated DEGs between Sed21% and Sed13% animals, indicating that Kupffer cells in Sed mice are highly sensitive to a moderate increase in fat calories (Fig. 7A, Supplementary Table 3). Upregulated genes were highly enriched in the biological process of “cell activation” and “phagocytosis”, suggesting an activated state of Kupffer cells in Sed21% mice (Fig. 7B). Upregulated genes were also associated with “regulation of interleukin-6”, specifically with the “negative regulation of IL-6 production”, suggesting an anti-inflammatory response of Kupffer cells in Sed21% mice. Upregulated genes were not or minimally enriched (FDR > 1E-5) in lipid and FA related metabolic processes, “regulation of lipid localization” being the most significantly enriched (Fig. 7B). In contrast, downregulated genes in Sed21% mice were highly enriched in lipid and carboxylic acid metabolic genes, both biosynthetic and catabolic, similar to the pattern of downregulated genes in hepatocytes (Fig. 5D). This response to the 21% diet is not surprising given that Kupffer cells are among the first in the liver exposed to material passing from the gastrointestinal tract via the portal circulation.
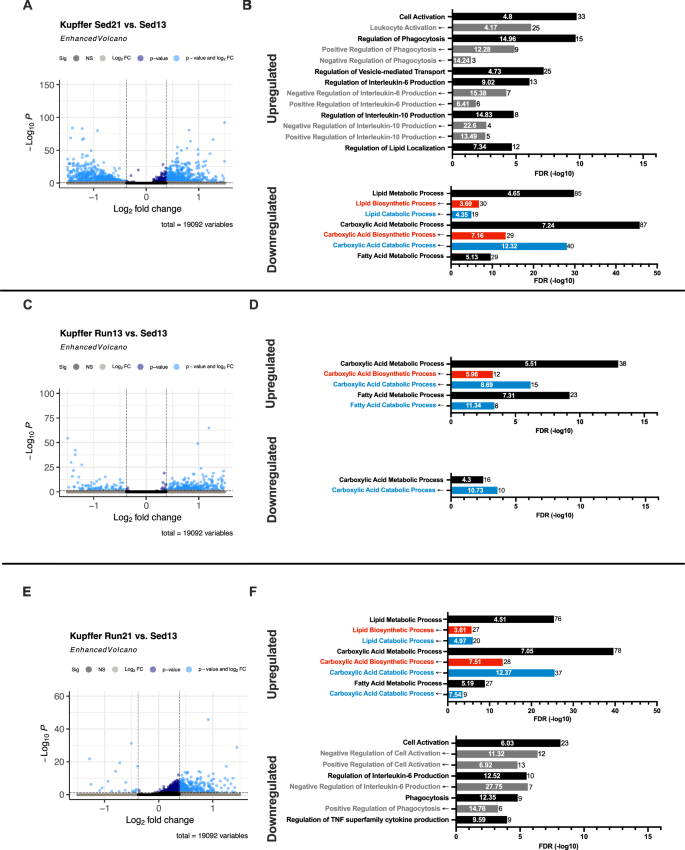
A 21% diet-induced change in gene expression in Sed males. Up- (+) and downregulated (–) genes are indicated by light blue dots in a volcano plot. B Enrichment of Sed21% vs. Sed13% DEGs in GO biological processes. Upregulated genes are enriched in immunity-related biological processes. C–F Same as (A, B) but for Run13% vs. Sed13% and Run21% vs. Sed13% DEGs, respectively.
The Run maternal environment produced 202 up- and 109 downregulated genes in the offspring on the 13% diet, and DEGs were associated with metabolic but not inflammatory processes, as expected. Upregulated genes were highly enriched in carboxylic acid and FA metabolic, predominantly catabolic processes (Fig. 7C, D, Supplementary Table 3), while downregulated genes were enriched in “carboxyl acid catabolic processes” but with a low enrichment score (Fig. 5G, H, Supplementary Table 3), suggesting a pattern promoting metabolic health. Overall, Kupffer cell transcriptomic changes to a 21% fat calorie diet in Sed mice and Run maternal environment were very different, unlike the transcriptomic changes in hepatocytes.
The 301 upregulated genes in Kupffer cells in Run mice on the 21% fat, in contrast to Sed21% mice, showed no sign of cell and immune activation and enriched biological annotations were purely limited to metabolic processes, especially carboxylic acid and FA catabolic processes, consistent with the spared liver phenotype of Run21% mice (Fig. 7E, F; Supplementary Table 3). However, the 107 downregulated genes were enriched in “phagocytosis”, particularly in “the positive regulation of phagocytosis suggesting an antiphagocytic effect. This, however, was balanced by the enrichment of downregulated genes in the ”negative regulation of cell activation and interleukin-6 production” annotations. Overall, the Kupffer cell transcriptome of Run21% mice seems to be the mirror image of the Sed21% transcriptome as 54% of the downregulated genes in the former are upregulated in the latter (enriched in phagocytosis and cell activation processes) and similarly, 76% of the upregulated in Run21% are downregulated in Sed21% (enriched in carboxylic acid and FA metabolic processes).
Finally, we analyzed differential gene expression in stellate cells, another non-parenchymal cell with star-like morphology and 3–4 arm-like projections52. We found 78 up- and 59 downregulated genes in Sed21% mice, relative to Sed13% mice (Supplementary Tables 3 and 4). Although upregulated genes were significantly enriched in only one biological process (FDR < 5E-2), “cell projection morphogenesis”, stellate cell projections have been implicated in cell-to-cell communication. Stellate cells are the main effectors in liver fibrosis due to their capacity to transdifferentiate into collagen-producing myofibroblasts53,54, but enrichment in “collagen fibril organization” driven by the collagen genes Col3a1 and Col14a1 was not significant after correction for multiple testing. Downregulated genes were enriched in carboxylic acid metabolic processes, again biased to catabolic processes, like downregulated genes in hepatocytes and Kupffer cells. Overall, the stellate transcriptome indicated no activation or fibrosis consistent with the moderate non-alcoholic fatty liver disease (NAFLD)-like liver pathology of Sed21% mice.
The running maternal environment resulted in 211 upregulated genes that were again enriched in carboxylic acid and FA metabolic processes, in particular catabolic processes, supporting a beneficial effect (Supplementary Tables 3 and 4). The 78 downregulated genes were not enriched in any GO functional category.
We found only a few stellate cells DEGs in Run21% vs. Sed13% (9 up- and 17 downregulated (Supplementary Table 3) with no significant enrichment in any biological function, indicating no major effect of the 21% fat calorie diet on stellate cells of Run mice.
link