Anti-inflammatory effects of Bifidobacterium infantis M-63 during the early postnatal period in term infants
Study design and participants
The present study is a single-center, placebo-controlled, double-blinded randomized trial, which is an additional analysis of a randomized clinical trial designed to evaluate the effect of B. infantis M-63 on healthy-term infants.36 Briefly, the study recruited healthy women who were to give birth to a healthy, full-term baby between October 2019 and August 2021. Eligible babies were healthy newborns born between 37 and 42 weeks. Exclusion criteria for mothers were a diagnosis of serious liver, kidney, cardiovascular, respiratory, endocrine, metabolic, or psychiatric disease, or planned use of other probiotics. Infant exclusion criteria were multiple birth, small for gestational age (SGA), large for gestational age (LGA), significant medical complications, exposure to antibiotics, and if deemed inappropriate by the investigator. Primary outcomes assessed relative abundance of Bifidobacterium, predominant proportion, and whether there was an increase in the number of copies in feces. Secondary outcomes included effects on gastrointestinal tract tolerance, child health (e.g., crying time, fever), fecal pH, SCFAs, sIgA, and calprotectin.
Healthy-term infants were randomized to receive either 1 billion colony-forming units of B. infantis M-63 (B. infantis M-63 group) daily or sterilized dextrin (placebo group) from ≤7 days to 3 months of age. Demographic information was collected for each participant and their parents. In addition, mothers kept daily records, including the number of times they fed their infants breast milk and formula. The study was sufficiently explained to the parents before enrollment, and written informed consent was obtained from the parents of all study participants. The study was conducted in accordance with the Declaration of Helsinki and approved by the Research Ethics Committee of Matsumoto City Hospital on September 27, 2019. The study was registered in the UMIN Clinical Trials Registry (UMIN000038351) in advance.
Fecal sample collection
Two types of infant fecal samples were collected from the infants’ diapers by their parents using fecal collection tubes (Techno Suruga Laboratory Co., Ltd., Shizuoka, Japan): the first type were fresh samples before intake and at 1 month of age, and the second were those preserved with a guanidine thiocyanate solution before intake, 1 week after intervention, and at 1 month and 3 months of age. All the samples, collected at home, were transferred to the hospital and stored in −20 °C freezers. After being transported to the laboratory with dry ice, they were stored at −80 °C. Fresh fecal samples were used for SCFA, cytokine, and metabolite analyses, and fecal samples with preservative solution were used for 16S rRNA gene sequencing.
Fecal DNA extraction and microbial, SCFA, and biomarker analysis
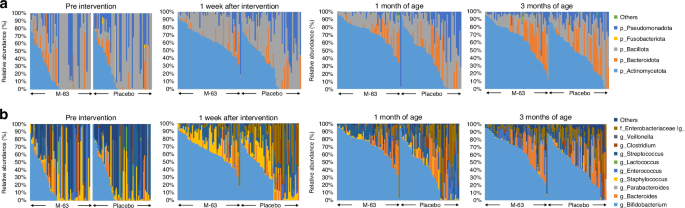
DNA extraction and microbial analysis were performed as described previously.36 The V3–V4 region of the bacterial 16S rRNA gene was amplified and sequenced on an Illumina MiSeq instrument (Illumina, San Diego, CA) for microbial analysis. Sequences aligned with GRCh38 and phiX reads were subsequently eliminated. The remaining sequences were analyzed using QIIME2 software version 2017.10. DADA2 was used to eliminate potential chimeric sequences and trim bases of the reads.45
Taxonomic classification was conducted using Greengenes 13.8 data, and alpha diversity was calculated with the QIIME2 software. Principal coordinate analysis (PCoA) and partitioning around medoid (PAM) clustering were conducted using R (version 4.3.0; R Foundation for Statistical Computing, Vienna, Austria). Enterotype clustering was performed at the genus level.39 Jensen–Shannon distance (JSD) and PAM clustering were applied for sample clustering, and the Calinski–Harabasz index was used to determine optimal enterotype clustering. Hierarchical clustering analysis was performed using the MeV suite version 4.9 ( with distances calculated based on the Pearson correlation of enterotype transition data. Fecal SCFAs (acetic acid, propionic acid, and butyric acid) were determined using gas chromatography, whereas secretory immunoglobulin A (sIgA) and calprotectin were measured using the corresponding enzyme-linked immunosorbent assay kit as previously described.36
Fecal cytokine analysis via multiplexed immunoassays
IL-1β and IL-8 levels were assessed using U-PLEX Biomarker Group 1 (human) Assays (Meso Scale Discovery, Rockville, MD). IFN-γ, IL-6, and TNF-α were measured with the S-PLEX Proinflammatory Panel 1 (human) Kit (Meso Scale Discovery). Additionally, IL-5 and IL-22 were quantified using the S-PLEX Human IL-5 and IL-22 Kits, respectively (Meso Scale Discovery), according to the manufacturer’s instructions.46
Briefly, 50–100 mg of fecal samples per participant were diluted 10-fold with a 100:1 mixed buffer of phosphate-buffered saline and proteinase inhibitor (Nacalai Tesque, Kyoto, Japan). Next, samples were agitated in a bead crusher (TAITEC, Saitama, Japan) with 5 mm stainless steel beads (QIAGEN, Valencia, CA) for three cycles of 2 min each. After centrifugation, the supernatant was collected, and each cytokine was measured using the respective kit. The standard and samples were assayed in duplicate using the Meso Scale Discovery (MSD) Quick Plex instrument and analyzed using the MSD software. Cytokine values were normalized to the fecal weight, and values below the detection limit were defined as zero.
Metabolite extraction
To each stool sample, 10–40 mg, 200 μL of methanol solution was added. The sample was crushed at 2000 rpm for 30 s with 300 mg of zirconia beads, followed by centrifugation at 10,000 × g at 4 °C for 10 min, and the supernatant was collected. The supernatant volume was filtered through a Nanosep 3 Komega microconcentrator (Pall, Port Washington, NY) and centrifuged at 9100 × g at 4 °C for 120 min. The filtrate was collected and treated with a centrifugal evaporator for 30–60 min until dryness. All pretreated samples were stored at −20 °C before measurement.
Quantification of fecal metabolite concentrations through LC-MS/MS
Fecal metabolites were determined using liquid chromatography–tandem mass spectrometry (LC-MS/MS) with a Vanquish high-performance liquid chromatograph coupled to TSQ-FORTIS (Thermo Fisher Scientific, Waltham, MA), as previously described.47 The analysis included 19 B. infantis M-63 and 23 placebo samples pre-ingestion, and 25 B. infantis M-63 and 16 placebo samples at 1 month. We measured indole-3-lactic acid (ILA), indole-3-aldehyde (IAld), indole-3-acetic acid (IAA), indole-3-propionic acid, 4-hydroxyphenyl-lactic acid (HPLA), 3-phenyl-lactic acid (PLA), tryptophan (Trp), tyrosine (Tyr), and phenylalanine (Phe) levels. A list of these compounds and their analytical conditions for selected reaction monitoring analysis is shown in Table S1.
LC-MS/MS quantification was performed using an XBridge® C8 column (4.6 × 150 mm, 3.5 μm) (Waters Corporation, Milford, MA). The mobile phases included water with 0.05% (v:v) formic acid (A) and methanol (B), with a flow rate of 0.2 mL/min. The gradient elution profile was as follows: (i) 2% B for 2 min, (ii) increase from 2% to 65% until 40 min, (iii) increase from 65% to 99% until 45 min, (iv) hold at 99% until 55 min, (vii) decrease from 99% to 2% until 60 min, and (viii) hold at 2% until 75 min. All chemical reagents used were of analytical grade.
Statistical analysis
Differences in microbiota between groups at the phylum and genus levels were assessed using ALDEx248; q < 0.05 was considered statistically significant. The Wilcoxon rank-sum test was used to compare the two groups, whereas Fisher’s exact test was used to analyze the distribution of infants within each enterotype or cluster. The Kruskal–Wallis test was used to confirm the presence of groups with the same distribution of cytokine levels, metabolites, and breastfeeding rate of infants in each enterotype, and Dunn’s multiple comparison test with Bonferroni correction was employed to identify differences within groups. Statistical analyses were conducted using R software (version 4.3.0) and SPSS software (version 28.0; IBM Corp., Armonk, NY); p < 0.05 was considered statistically significant.
link